Perez-Roldan, Juan and Henn, László and Bernues, Jordi and Torras-LLort, Monica and Tamirisa, Srividya and Belloc, Eulalia and Rodriguez-Munoz, Laura and Timinszky, Gyula and Jimenez, Gerardo and Mendez, Raul and Carbonell, Albert and Azorin, Fernando (2025) Maternal histone mRNAs are uniquely processed through polyadenylation in a Stem-Loop Binding Protein (SLBP) dependent manner. NUCLEIC ACIDS RESEARCH, 53 (7). No. gkaf288. ISSN 0305-1048
|
Text
gkaf288.pdf - Published Version Available under License Creative Commons Attribution Non-commercial. Download (10MB) | Preview |
|
![[img]](https://real.mtak.hu/235766/7.hassmallThumbnailVersion/gkaf288figgra1.jpeg)
|
Text
gkaf288figgra1.jpeg - Published Version Available under License Creative Commons Attribution Non-commercial. Download (296kB) | Preview |
Abstract
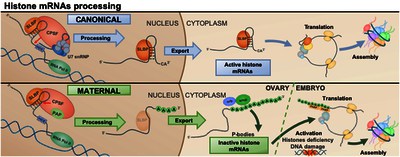
During early embryogenesis the zygotic genome remains transcriptionally silent and expression relies on maternally deposited products. Maternal deposition of histones is crucial to preserve chromatin integrity during early embryo development, when the number of nuclei exponentially increases in the absence of zygotic expression. In the Drosophila embryo, histones are maternally deposited as both proteins and mRNAs. Histone transcripts are the only nonpolyadenylated cellular mRNAs. They contain a highly conserved 3 ' UTR stem-loop structure, which is recognized by the Stem-Loop Binding Protein (SLBP) that, in conjunction with U7 snRNP, regulates their unique 3 '-end processing. Here we report that, unexpectedly, maternal histone mRNAs are polyadenylated and have a truncated 3 ' stem-loop. This noncanonical 3 '-end processing of maternal histone mRNAs occurs at their synthesis during oogenesis and requires SLBP, but not U7 snRNP. We show that maternal histone transcripts are subjected to cytoplasmic poly(A) tail elongation by Wisp, which results in their stabilization and is a requisite for translation. We also show that maternal histone transcripts remain largely quiescent and that their translation is activated upon loss of the embryonic linker histone dBigH1, which impairs chromatin assembly and induces DNA damage. Here, we discuss possible models to integrate these observations.
| Item Type: | Article |
|---|---|
| Additional Information: | Funding Agency and Grant Number: MICIN/AEI [PGC2018-094538-B100, PID2020-119248GB-I00, PID2021-123303NB-I00]; FEDER, una manera de hacer Europa; Generalitat de Catalunya [SGR2017-475] Funding text: This work was financed by grants PGC2018-094538-B100, PID2020-119248GB-I00, and PID2021-123303NB-I00 from MICIN/AEI 10.13039/501100011033 and "FEDER, una manera de hacer Europa", and of the Generalitat de Catalunya (SGR2017-475). |
| Uncontrolled Keywords: | EXPRESSION; ACTIVATION; OOGENESIS; translational control; poly(A) polymerase; 3' END; Cytoplasmic polyadenylation; DROSOPHILA ORB GENE; FACTOR CPSF-73; |
| Subjects: | Q Science / természettudomány > QH Natural history / természetrajz > QH301 Biology / biológia |
| SWORD Depositor: | MTMT SWORD |
| Depositing User: | MTMT SWORD |
| Date Deposited: | 18 Mar 2026 15:56 |
| Last Modified: | 18 Mar 2026 15:56 |
| URI: | https://real.mtak.hu/id/eprint/235766 |
Actions (login required)
 |
Edit Item |